News
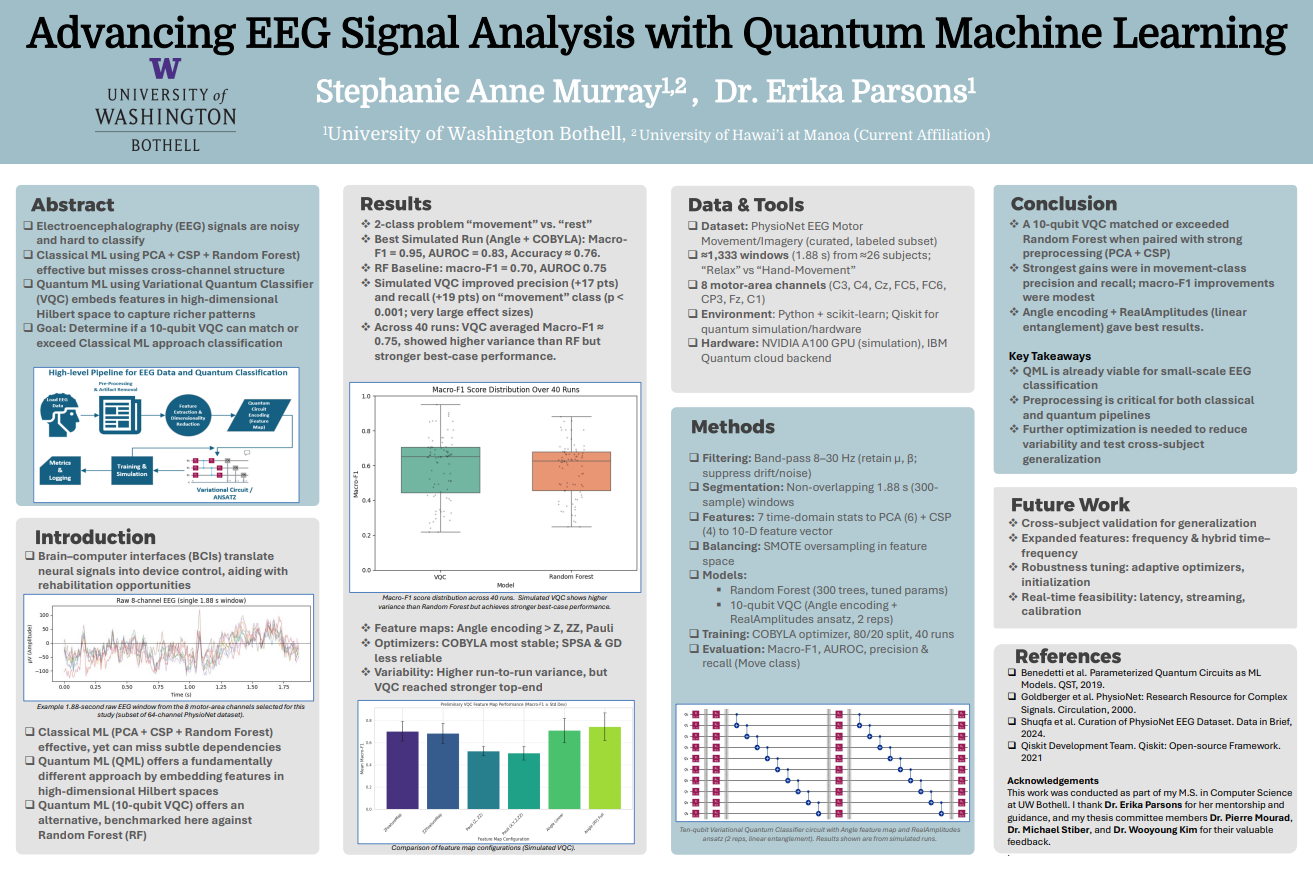
November 2025 — SC25 poster presentation
I presented my master’s thesis poster, Advancing EEG Signal Analysis with Quantum Machine Learning, at Supercomputing 2025 (SC25) in St. Louis. The poster summarized results from my written thesis comparing variational quantum circuits to a tuned Random Forest baseline.
December 2025 - Michael Bruno Award for Excellence in Microbiome Science
Received the Michael Bruno Award for Excellence in Microbiome Science through the C-MĀIKI #MahiMicrobe2025 program for a proposed Rapid ‘Ōhi‘a Death microbiome project focused on comparing microbial communities from surviving and infected trees using amplicon sequencing, computational biology, and machine learning, in collaboration with Dr. Lenore Pipes.